Examples of using CRISPRs Web service
Summary
Example 1 : Find CRISPRs in Aquifex aeolicus VF5
General overview
Data used for this sample is a fasta format file of the chromosome circular of Aquifex aeolicus VF5 (RefSeq=NC_000918, GenBank id=AE000657).
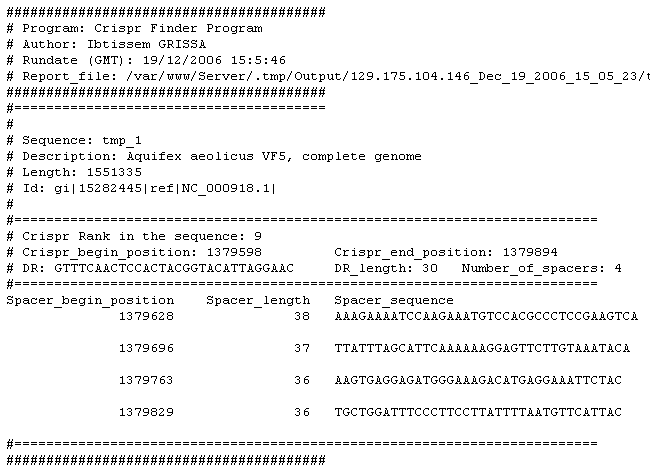
After quering this sequence by CRISPR Finder, the list of detected CRISPRs is given :
Questionable CRISPRs are non confirmed CRISPRs because they are short clusters (less than 3 spacers).
For each cluster, there is a link for more details.
CRISPRs candidates localization links to a figure showing the different clusters positions in the circular genome.
CRISPR displaying
The list of clusters is given with detailed information about their position, size, number of repeats,...
A colour schema is associated to Direct Repeats (DR) and spacers (see CRISPR definition). DRs are yellow whereas spacers have different colours to show that they are different.
Upload output
There is also a possibility to upload the summary of CRISPR properties as shown below :
or to get the list of spacers in Fasta format to use for further analyses by clicking on the Display spacers button :
Once CRISPRs are detected, there are obviously further analyses to do. In order to help users in this task, we offer a list of utilities available on the left menu :
- BLAST spacers
- Get Flanking sequences
- Identical DRs
BLAST spacers
The list of the relative spacers is given and spacers of interest should be selected :
BLAST searches are done against the Genbank databases. Matches are considered to be significant if the matching sequence size is about 70% the queried spacer size and the E-value is less than 0.1.
Get Flanking sequences
This is a flexible tool to obtain the flanking sequences and the CRISPR sequence with the possibility of modifying their respective positions.
Getting the flanking sequences will be useful especially to define the leader sequence.
Identical DRs
Comparing a CRISPR to known CRISPRs is possible by the comparison of Direct Repeats. This tool is useful especially to confirm a questionable CRISPR or to get an idea about horizontal transfer of the structure.
Only identical DRs are shown with a reference to related specie.